Journal: bioRxiv
Article Title: Synthetic genomic reconstitution reveals principles of mammalian Hox cluster regulation
doi: 10.1101/2021.07.07.451065
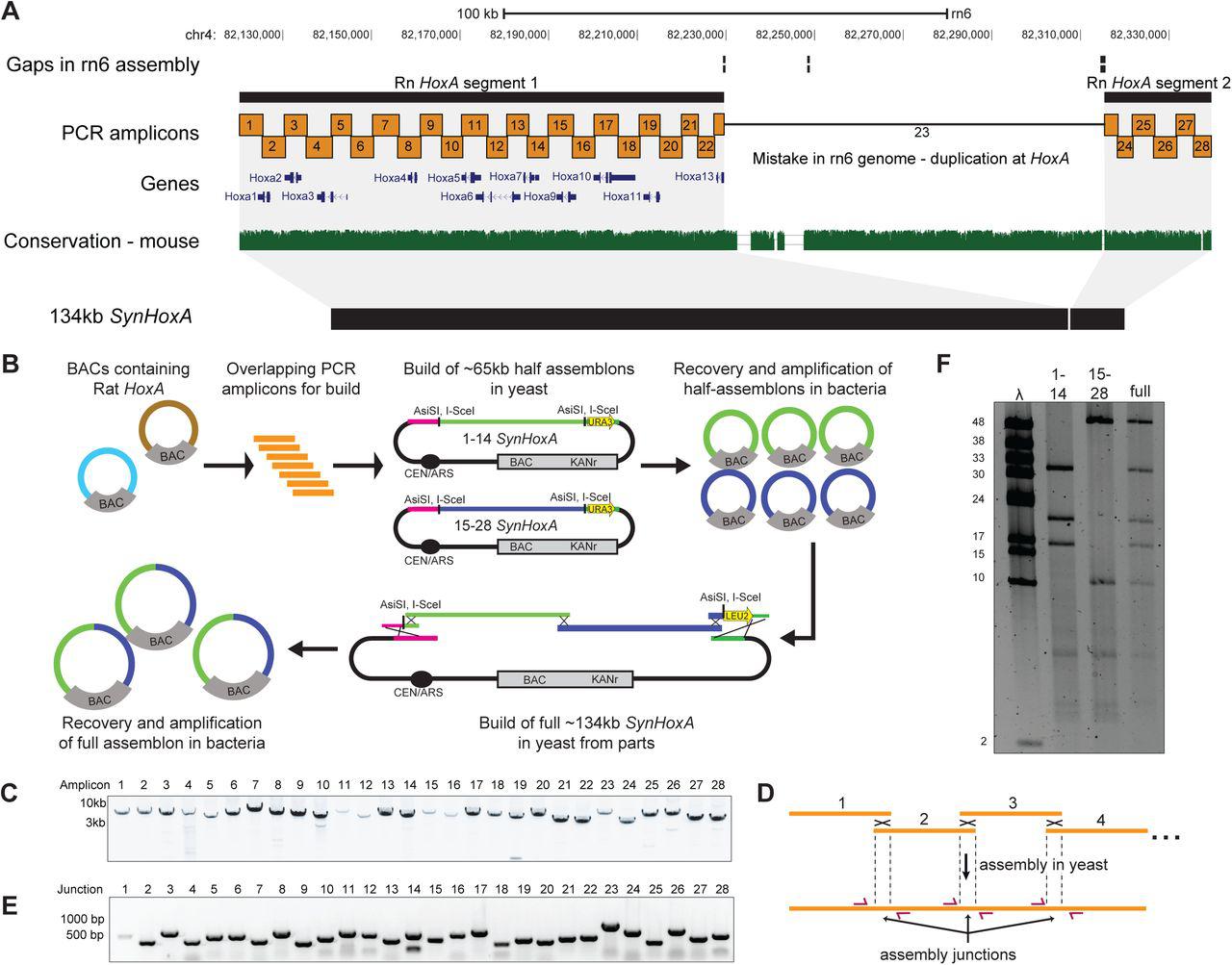
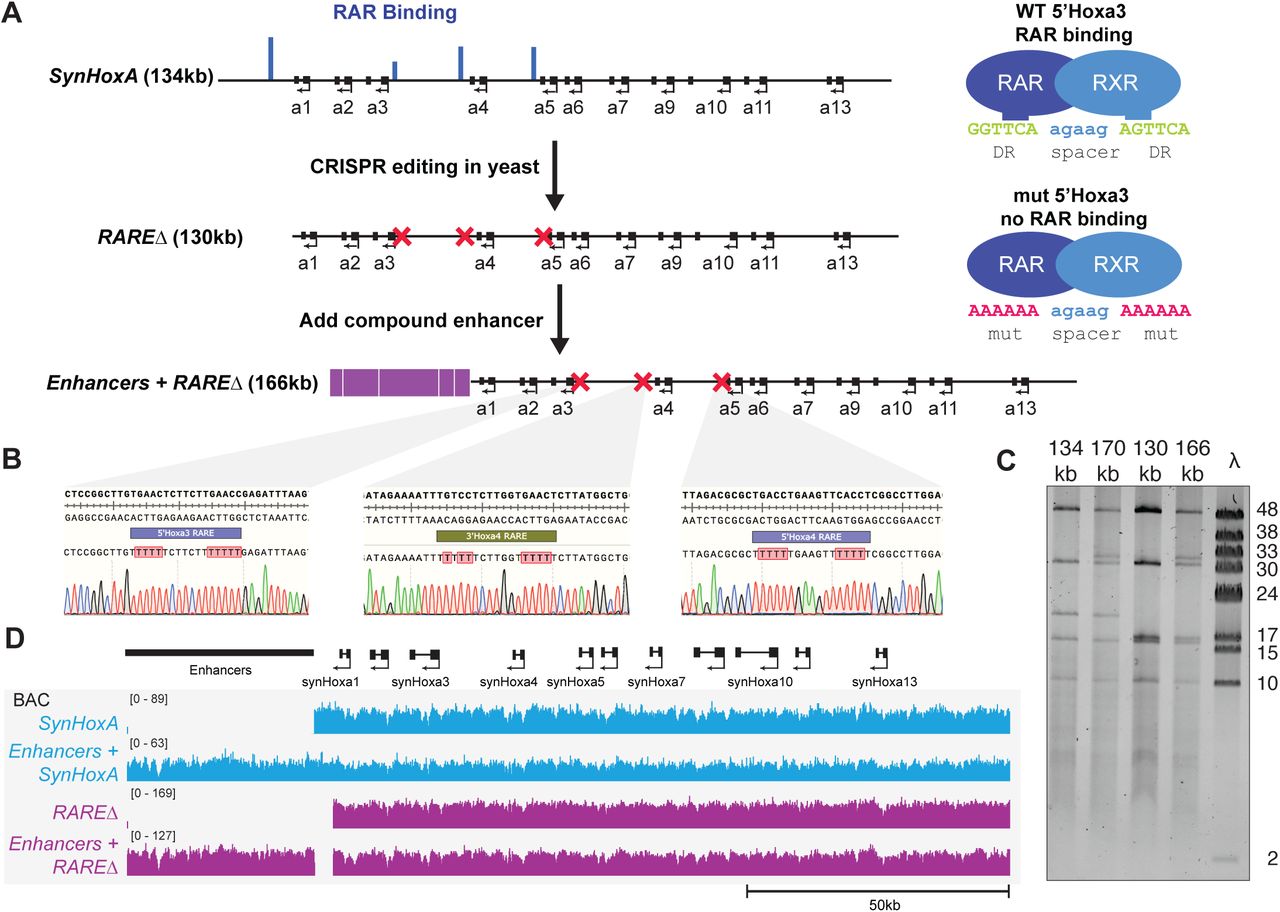
Figure Lengend Snippet: (A) Layout of rat HoxA locus in the rn6 genome assembly. The rn6 genome includes an erroneous duplication at the HoxA locus between gaps in the assembly. The SynHoxA assemblon sequence is based on bringing together the two ‘separate’ RnHoxA segments. The sequence was segmented into 28 ∼5kb PCR amplicons with terminal homology of ∼200bp to adjacent amplicons. Conservation to the mouse genome is depicted using the multiz track from the UCSC genome browser. (B) Schematic depicting the assembly workflow for the 134kb SynHoxA assemblon. BACs containing Rat HoxA were used as PCR template to generate 28 segments tiling the entire HoxA locus. These segments were co-transformed into yeast with appropriate linkers and assembly vector to build two ∼65kb half assemblons into centromeric yeast-bacteria shuttle vectors. These half assemblons are recovered to bacteria and amplified. Full 134kb assemblon was built from half assemblons after releasing them from the vector using terminal restriction enzymes ( AsiSI ) and transforming into yeast. Full assemblon was then recovered from yeast into bacteria for amplification and verification. (C) Agarose gel of the 28 PCR amplicons that tile the 134kb SynHoxA assemblon. (D) Strategy to PCR-screen yeast colonies derived from assembly experiments. Primers (red arrows) span assembly junctions and test presence/absence of amplicons in many yeast colonies. Reproduced from ref with permission from authors. (E) Agarose gel showing one yeast colony carrying the full 134kb SynHoxA assemblon verified manually for the presence of all assembly junctions, using the strategy outlined in panel D. (F) Half and Full 134kb SynHoxA assemblon BACs purified from E.coli were digested with PvuI and separated using field inversion gel electrophoresis (FIGE). Lambda monocut ladder sizes are indicated in kb. Band sizes correspond to expected fragments.
Article Snippet: 500 ng of lambda monocut ladder (New England Biolabs N3019S) were used as a molecular weight standard.
Techniques: Sequencing, Transformation Assay, Plasmid Preparation, Amplification, Agarose Gel Electrophoresis, Derivative Assay, Purification, Nucleic Acid Electrophoresis